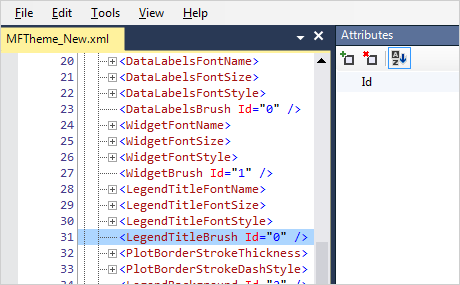
⌘ Command + Mouse Wheel or ⇧ Shift + ⌃ Ctrl + Mouse Wheel Rotate (pan and tilt) around the point where the mouse was clicked. The following table shows the available navigation commands using the mouse: Left Drag anywhere on the canvas to rotate (reslice) the image.Use ⌃ Ctrl + ⇧ Shift + Mouse Wheel, or ↑ Up and ↓ Down keys to zoom in and out.Right Drag anywhere on the canvas to translate the image.
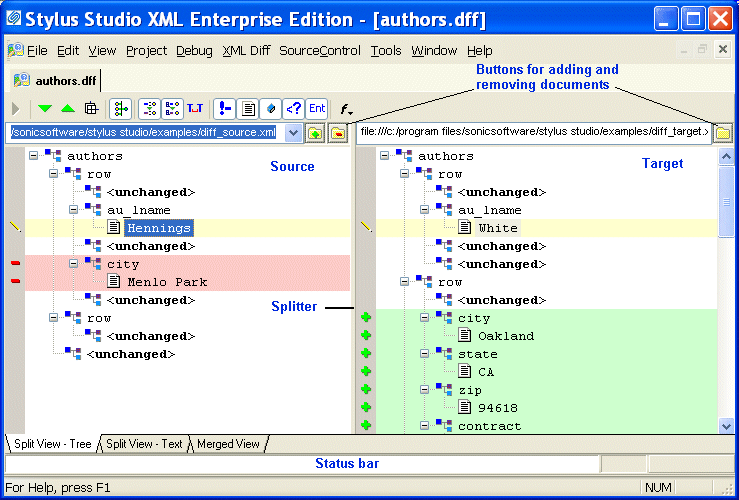
Use the mouse-wheel or keys to scroll through z slices.

You can browse the stack using the keyboard or the mouse. On startup, the middle slice of the first source (angle) is shown. Basic NavigationĪssuming you downloaded the drosophila melanogaster example dataset, you should see something like this: Please note, that support for the Imaris format is still experimental. ims files select Plugins › BigDataViewer › Open Imaris (experimental) from the Fiji menu. Imaris (Bitplane) uses a hierarchical data format (similar to BigDataViewer’s XML/HDF5 format). Subsequently select Plugins › BigDataViewer › Open Current Image which will launch the BigDataViewer with the sample image, for navigation by arbitrary re-slicing. Since Fiji relies of LOCI Bioformats library that means essentially all know microscopy file formats.įor example, open the sample image File › Open Samples › Mitosis (26MB, 5D stack) from the Fiji menu.
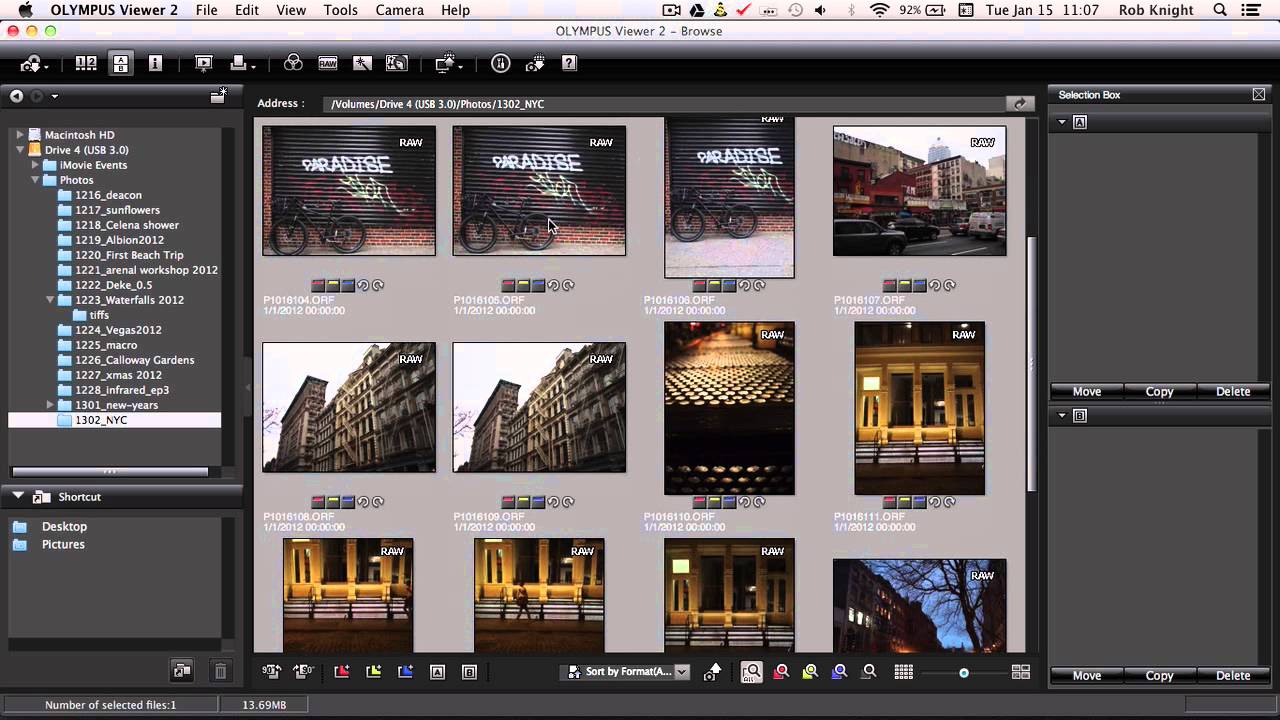
The BigDataViewer can be used to visualise and navigate any image that can be opened in Fiji. To start BigDataViewer, select Plugins › BigDataViewer › Open XML/HDF5 from the Fiji menu.
XML FILE VIEWER IMAGE DOWNLOAD
Download both the XML and the HDF5 file and place them somewhere next to each other.Īlternatively, you can create a dataset by exporting your own data as described below. This is an excerpt of a 6 angle 715 time-point sequence of drosophila melanogaster embryonal development, imaged with a Zeiss Lightsheet Z.1. You can download a small dataset from here, comprising two views and three time-points. Our custom XML/HDF5 is a special purpose hierarchical data format that optimises access to any part of large-multi view datasets using ImgLib2. There are various options: Multi-view data converted to XML/HDF5Ī special purpose of BigDataViewer is to visualise multi-view light sheet microscopy datasets. To use the BigDataViewer we need some example dataset to browse. You should have a sub-menu Plugins › BigDataViewer.
XML FILE VIEWER IMAGE REGISTRATION
The XML file contains metadata, for example the registration of sources to the global coordinate system. Images are represented as tiled multi-resolution pyramids, and stored in HDF5 chunked multi-dimensional arrays. The file format is based on XML and HDF5. This permits browsing to any location within a multi-terabyte recording in a fraction of a second. In a multi-angle, multi-channel SPIM sequence, each channel of each angle is a source.īigDataViewer comes with a custom data format that is is optimized for fast random access to very large data sets. For example, in a multi-angle SPIM sequence, each angle is a source. Each source provides one 3D image (for each time-point in the case of a time-lapse sequence). BigDataViewer was developed with multi-view light-sheet microscopy data in mind and integrates well with Fiji’s SPIMage processing pipeline.Ĭonceptually, the visualized data comprises multiple data sources. The BigDataViewer is a re-slicing browser for terabyte-sized multi-view image sequences.
XML FILE VIEWER IMAGE HOW TO
If you’d like to help, check out the how to help guide! Description The content of this page has not been vetted since shifting away from MediaWiki.
